We aim to create new molecular therapies for untreatable neurological diseases, ranging from epilepsy and autism to brain cancer and degeneration. By combining synthetic biology with neurodevelopmental research, we identify and target novel mechanistic interventions.
To achieve these aims, our team develops synthetic biology tools and applies them to the developing brain of mice and rats. A big focus is developing new CRISPR tools able to knock in precise genome edits directly in the developing bran, often by in utero electroporation, to track and manipulate neural circuits and their molecules as the brain develops. With these approaches and the generous support below, we are investigating the wiring patterns that underlie conditions on the neurodevelopmental spectrum, including, epilepsy, autism, and schizophrenia.
Let’s get into some specifics:
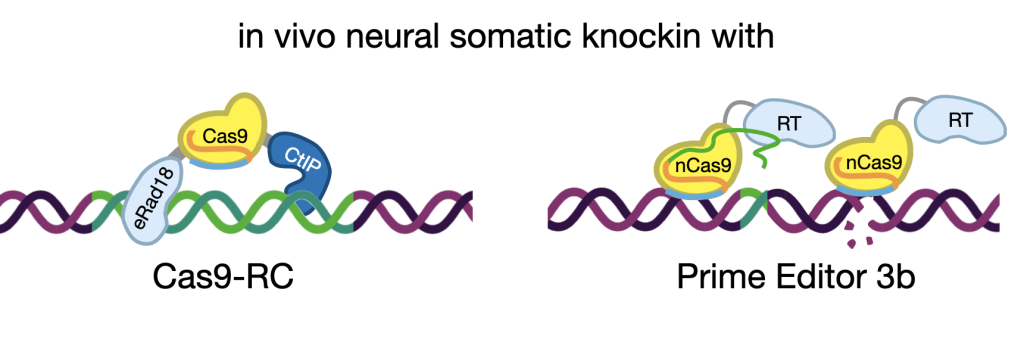
“Crispatics” somatic cell genome editing: in vivo knockin with Cas9-RC and Prime editors
We are in constant need of better tools. For instance, we would like a better way to edit the genome directly in relevant cells in situ, both for experimental purposes, and for future therapeutics. We’ve been tinkering with Cas9 to make it more precise for use in vivo, and came up with a new double-fusion we call “Cas9-RC” that is made for knockin use in the brain (Richardson et al. CRISPR J, 2023). We have been using in vivo Prime Editing to model individual patients with epilepsy in wild-type mice as personalized avatars for exploratory pharmacotherapy (Robertson et al. bioRxiv, 2024), the first of its kind! Finally we combine focus ultrasound and nanoparticle technologies to non-invasively deliver editors to the brain (Kwak et al. ACS Nano 2024), paving the way for future genome therapeutics!
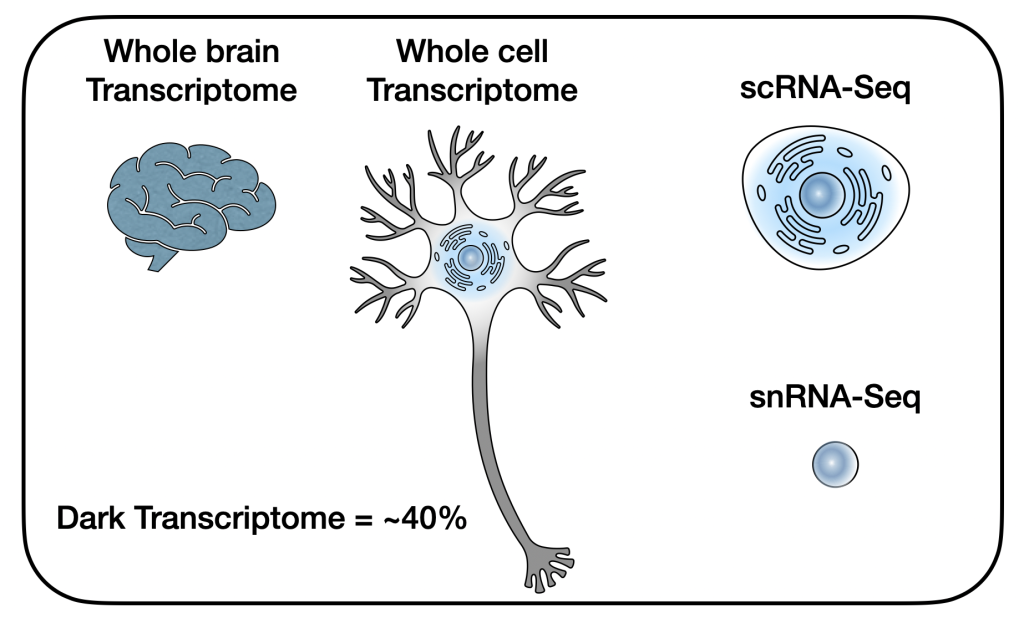
Dark Transcriptomes and mTOR Outposts
Neurons can be enormous cells, with much of their molecular diversity far away from their cell body. It turns out that the almost 40% of all transcripts in the brain are in cellular processes that get left behind by single-cell sequencing approaches (Ament & Poulopoulos, Curr Opin Neurobiol, 2022). This “dark transcriptome” remains unseen in scRNAseq datasets, and represents a mysterious side of cell biology that is unique to neurons and glia. We are very interested to discover what happens in those processes, far away from the nucleus, where a lot of molecularly diverse and locally dynamic subcellular biology takes place that gives the brain its hallmark complexity.
A key molecule whose role we believe is to coordinate between the dark transcriptome and the cell body transcriptome is mTOR. Through subcellular structures we call “mTOR outposts“, we think the cell body can chose to respond to molecular conditions in the most distal reaches of neuronal processes. We have seen these structures in vitro and hypothesize exist in vivo, but we still don’t know quite what they do and how they affect cellular growth, circuit wiring, and synaptic plasticity; but we have some ideas! Read all about neuronal mTOR outposts in our hypothesis and theory article Altas et al. Front Cell Neurosci, 2022.
Circuit development
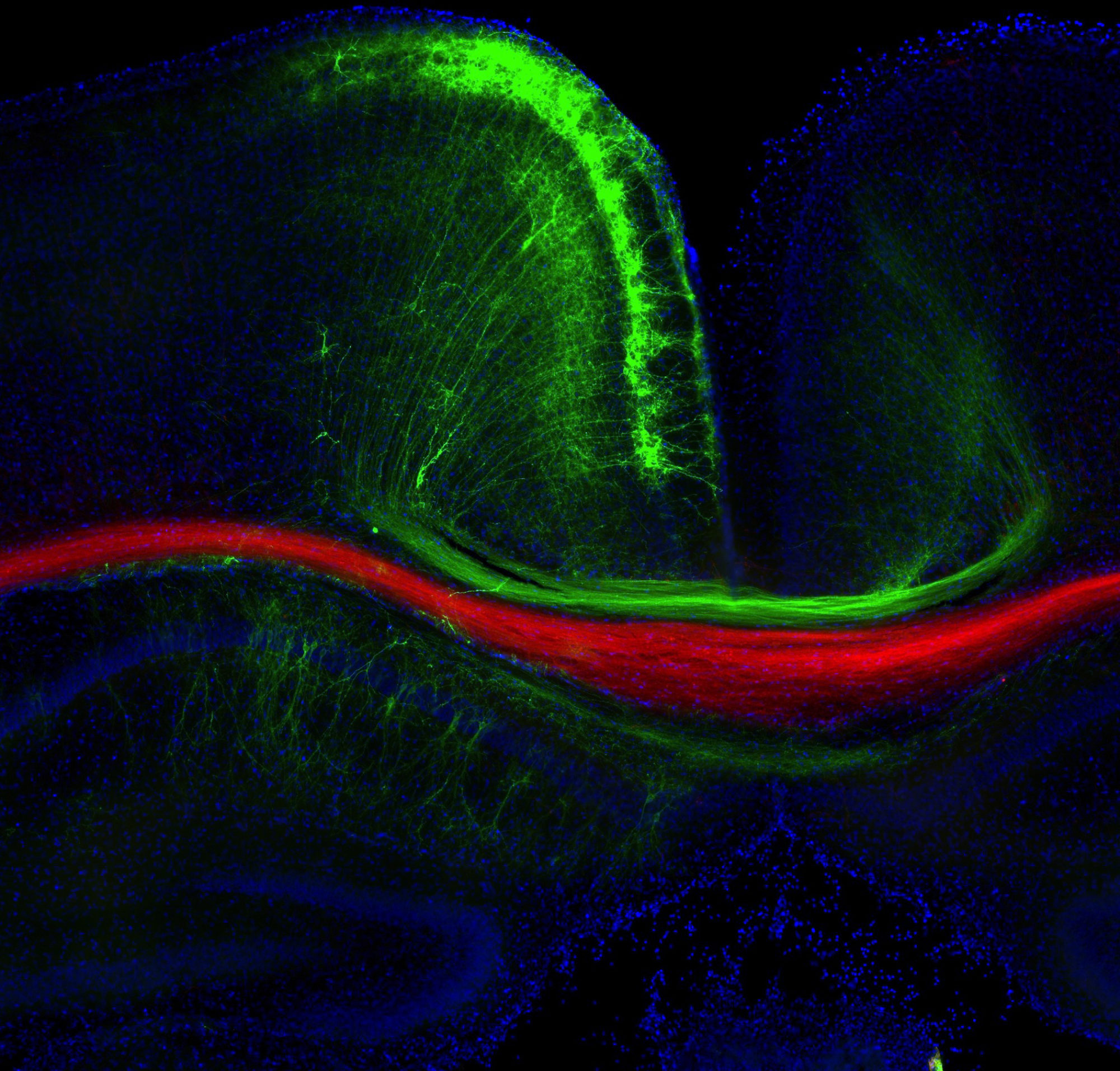
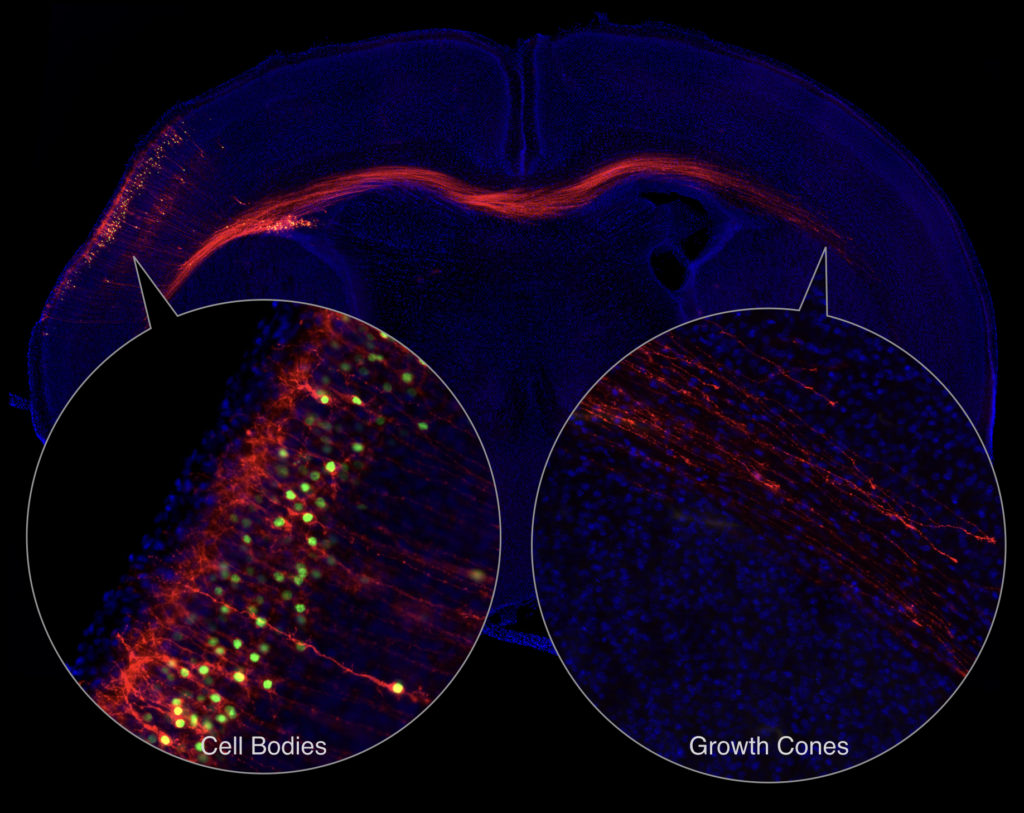
We previously developed an approach to identify local protein and RNA networks in axons that are in the process of forming connections in the developing brain (Poulopoulos et al, Nature, 2019). The approach combines in utero electroporation, subcellular fractionation, small-particle sorting, mass-spec and RNA-seq in order to reveal and quantify molecules that are in the farthest reaches of developing neurons.
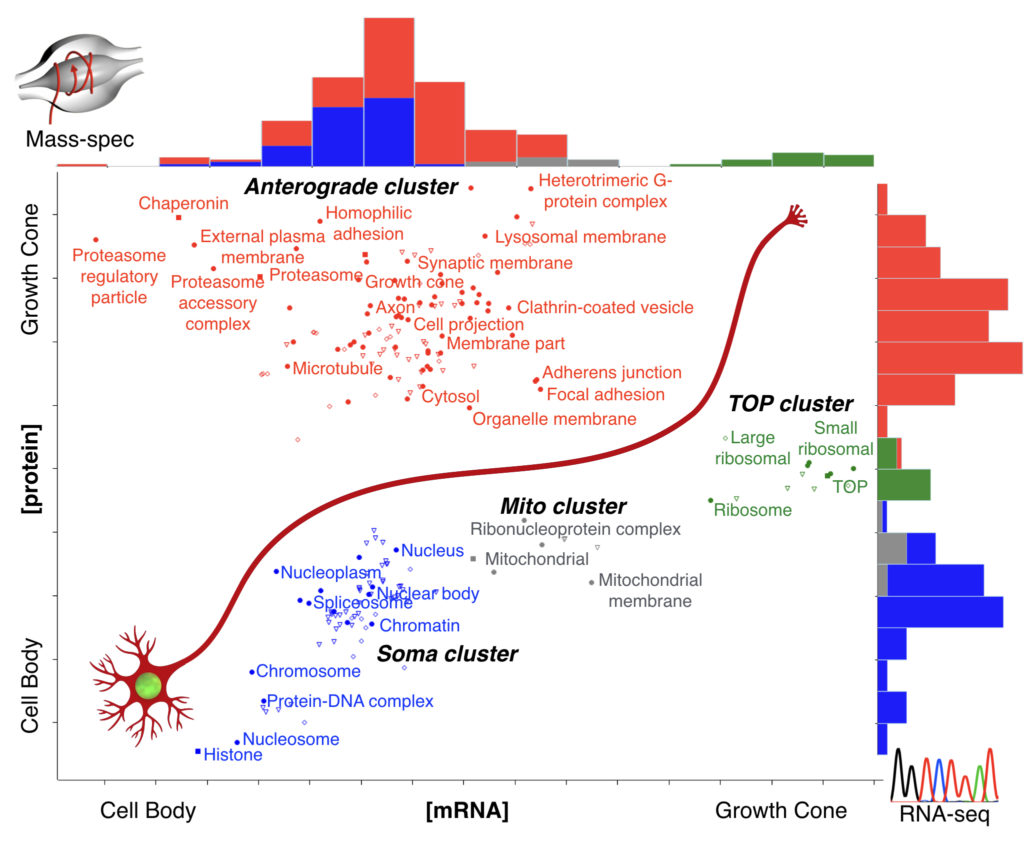
With this approach, we know which molecules concentrate in axons of a particular connection, giving us a “parts list” of the machinery that is implementing connectivity in vivo. Some of the biological processes we are currently following are cell adhesion, mTOR signaling, and local translation.
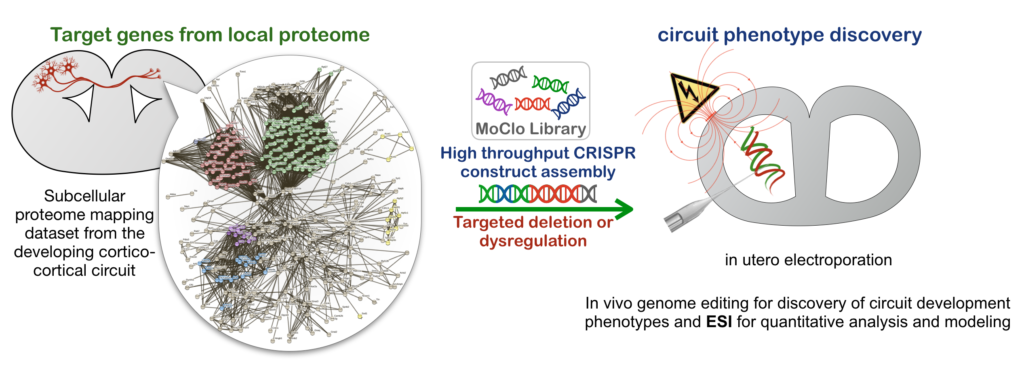
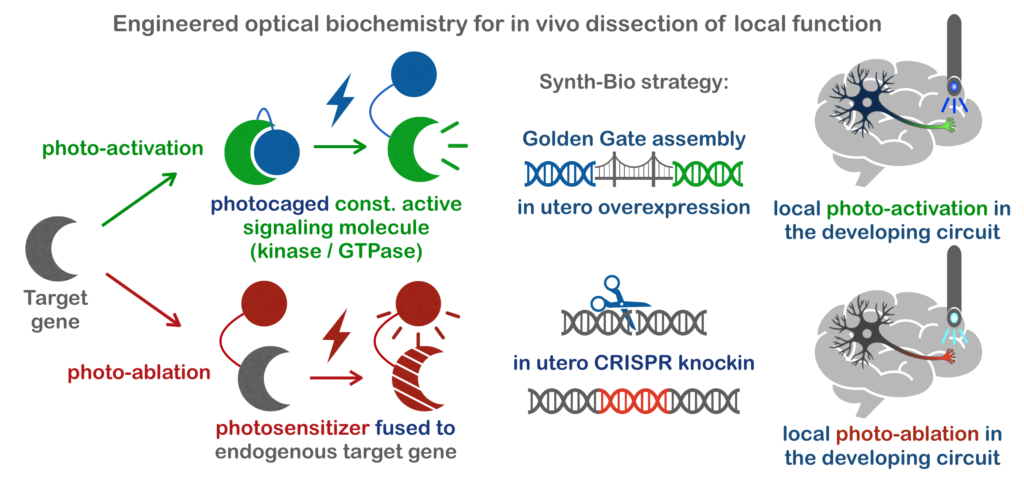
With this knowledge, we are targeting genes in these pathways using in vivo CRISPR genome editing for knockout and knockin. This allows us to screen for area-specific miswiring phenotypes, as well as to observe these molecules in vivo. With some added fancy footwork in the realm of synthetic biology, we can also functionally hijack the these molecules locally using light as they go through the steps of forming or modifying brain circuitry!
Synapse development
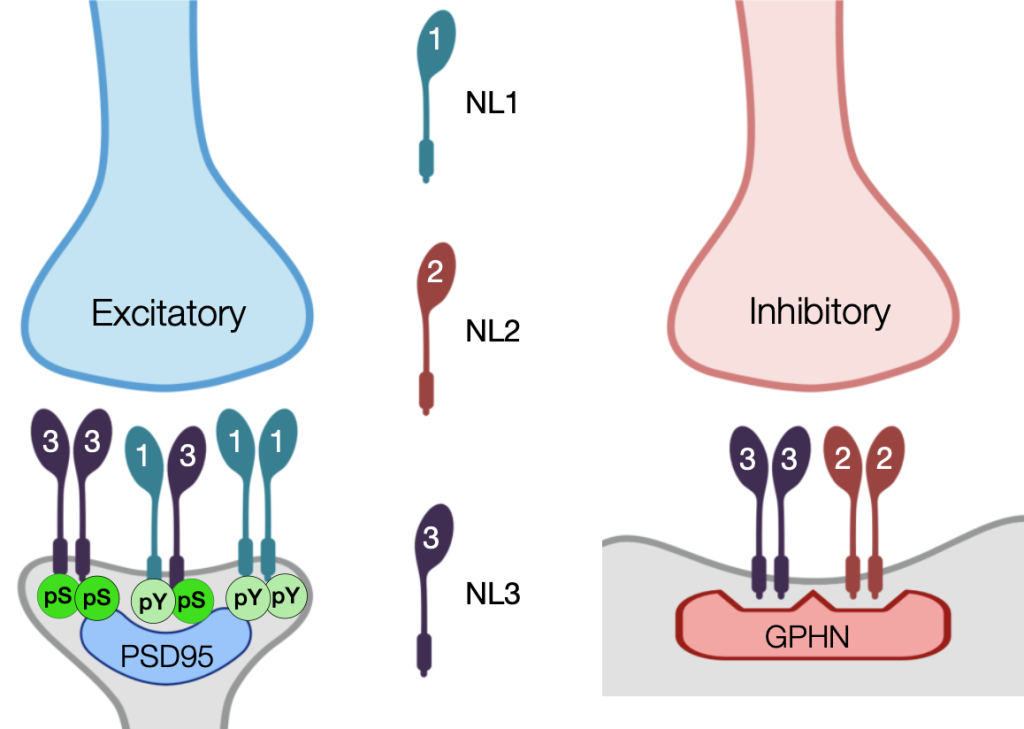
Beyond the formation of long-range connectivity, we look up close at the critical moment when a synapse develops to complete the connection between two neurons. We have a long-term interest in molecules like Neuroligins that coordinate processes across the synapse (Poulopoulos et al, Neuron, 2009; Poulopoulos et al, Biochem J., 2012; Altas et al, Biol Psychiatry, 2024). With our collaborators in the Blanpied lab across the street -the gurus of synapse nanostructure- we use in vivo CRISPR to tag endogenous synaptic proteins and examine their structure and function in the pre- and postsynaptic apparatuses. And with our collaborators at the Neurosurgery Translational Therapeutics Research Group, we harness neurodevelopment into therapeutics for brain cancer.
Mental illness and pathologies of brain wiring
We use in utero electroporation, in vivo genome editing, and 3D imaging to uncover the effects on brain circuitry of genetic variants associated with neurodevelopmental disorders and Mental illness. By introducing these variants using in vivo CRISPR in select circuits of the rodent brain, we aim to identify how these connections are altered to discover circuit underpinnings of conditions such as epilepsy, autism, and schizophrenia.
For these clinically relevant projects, we work closely with collaborating labs on campus. On our epilepsy projects we work with the lab of Peter Crino from the department of Neurology to create a full workflow from molecular, cellular, and electrophysiological endophenotyping, to EEG and behavioral monitoring in order to analyze clinically relevant mutations in mice .
On Schizophrenia, we collaborate with groups at the Maryland Psychiatric Research Center and Seth Ament at the Institute for Genome Sciences, who provide invaluable human brain imaging and genome data from cohorts of patients, which we get to test in the mouse cerebral cortex. On neurodevelopmental disorders we collaborate with the groups of Saima Riazuddin and Zubair Ahmed who have used human genetics of isolated populations to discovered a host of new recessive gene mutations leading to sever disorders of brain development, which we can examine molecularly in the embryonic mouse brain.
To go beyond descriptions of circuit aberrations, we have developed ESI, the Endophenotype Severity Index, a ratiometric value that allows generalizable quantification and modeling of circuit alterations associated with disease. With an interest in the circuitry of the cerebral cortex, with ESI we hope to document specific circuit and network defects in mouse models of mental illness, such as schizophrenia.
Hodaptics: cell-cell interactions in intact systems
The biology of cell-cell interactions is a beautifully complex and critical part of the development and function of multicellular systems. In the nervous system, this biology manifests in its extreme, with a given neuron directly contacting a bewildering and perpetually-changing number of direct cellular neighbors at any given time. It is this multitude and dynamics of cell-cell interactions that give nervous tissue its unique functionalities. However, the biology of cell-cell interactions is particularly difficult to study, as it requires experimenting on intact systems, while being able to detect and influence elusive transient interactions. To allow experimentation into this interesting biology, we have developed and are currently optimizing a new genetic technology, termed “hodaptics” (from Greek hodos- road, and haptein -to touch), that will allow us to detect and manipulate transient cell-cell interactions as they occur within intact systems such as the developing and learning brain, as well as other complex multicellular systems.