Axon bubbles: hidden structures in the developing brain, seen for the first time!
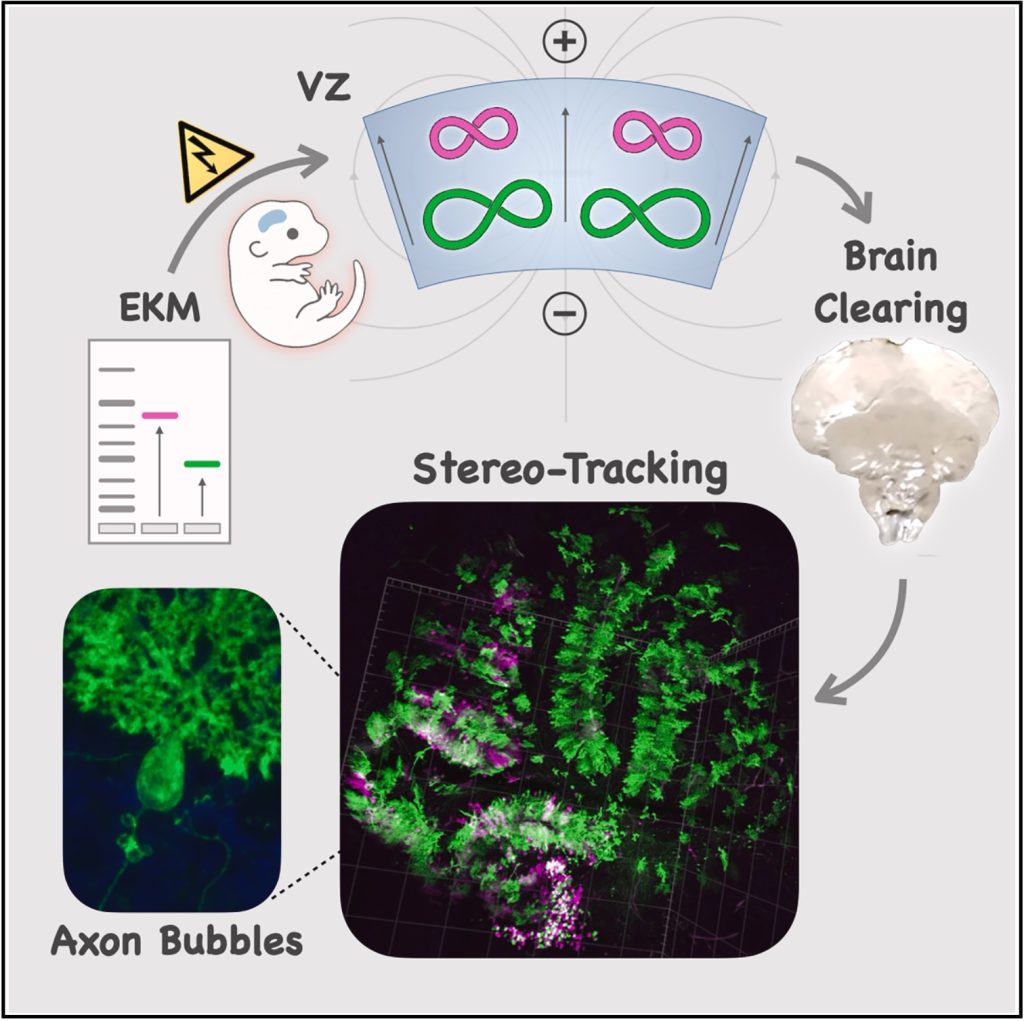
A new method we call Stereo-Tracking takes advantage of a bioelectrical interaction between DNA and cells in the embryonic brain to label the brain’s progenitors in 3D! Combined with brain clearing and light sheet imaging, it lets us track how different neural stem cell niches map onto brain circuits in ways we could not before.
Applying it to the cerebellum, the “little brain” at the back of the head, we stumbled upon a previously unknown cellular structure. We noticed bizarre membranous spheres clustering around the base of growing axons in the second postnatal week of development. They appear completely empty inside, making them invisible to previous labeling methods.
We named these mysterious structures “electrolucent axon bubbles“. Fun fact, we first confirmed their existence on the day of the spring equinox, when they appeared as radiant disks on electron micrographs; hence electrolucent, a small dedication to Apollo, the sun god 🌞. True science anecdote!
These bubbles form on Purkinje cell axons as granule cells squeeze past them during the largest neuronal migration in the brain (don’t forget: the cerebellum contains many more neurons than the rest of the brain combined)!
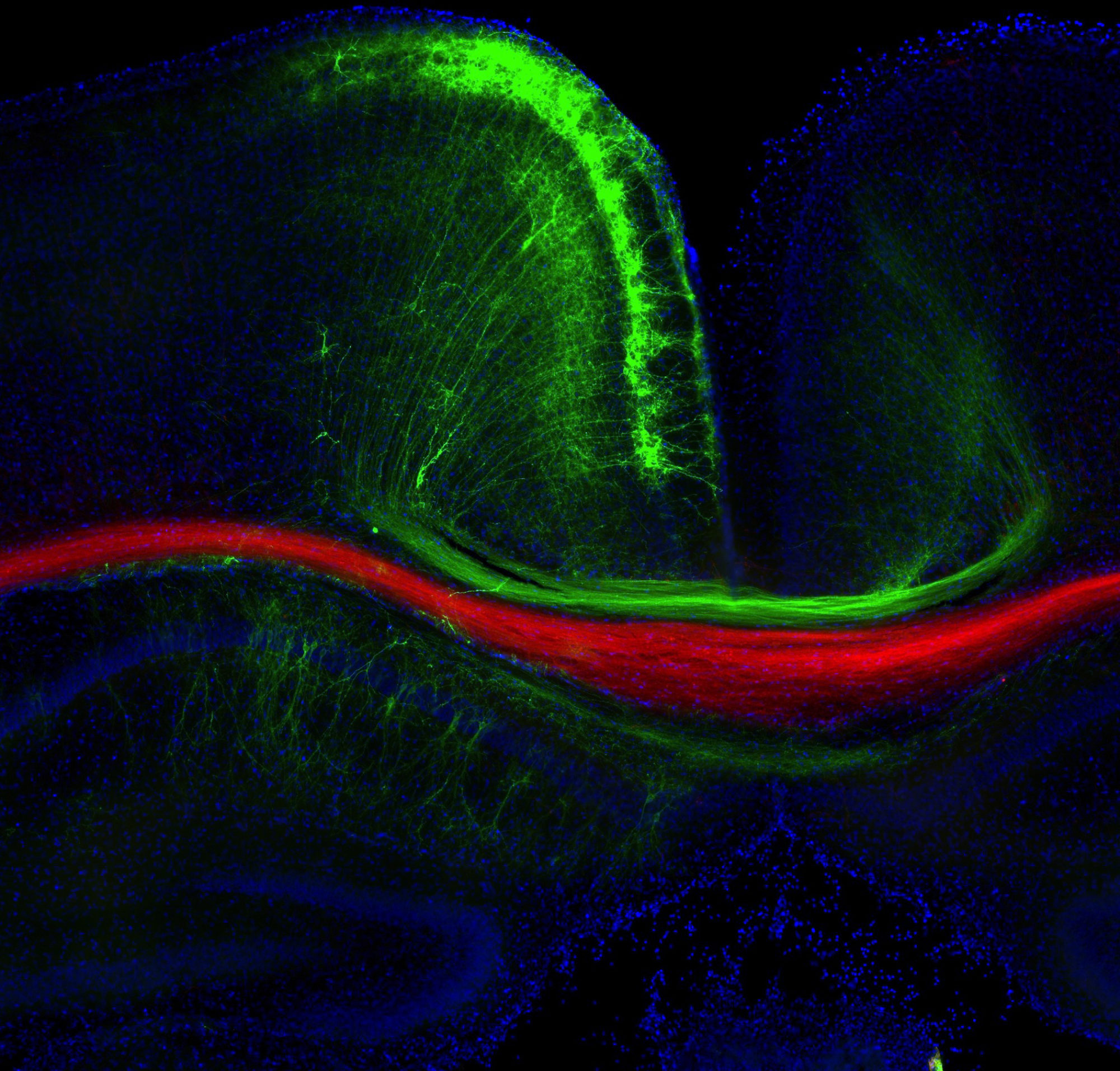
The light sheet rendering below shows Stereo-Tracked Purkinjes as they wire the developing cerebellum. Their striking circuitry quietly handles enormous computational workloads, like GPU arrays running alongside the cerebral cortex’s CPU. Kudos to Dr. Cheryl Brandenburg for these beautiful data!
Read the full story in our paper just out in Cell Reports Methods.
Huge congratulations to lead author Cheryl Brandenburg, the entire PouLab team at UMSOM, and our fantastic collaborator teams Roy Sillitoe at Baylor, Ben Cooper at the Max Planck Institute for Multidisciplinary Sciences, and Izumi Sugihara at Tokyo Medical and Dental University!
Big thanks as well to the NIH Common Fund, the Autism Research Institute, and the MPRC Training Program. Without your support, none of this would have been possible!